Scroll to:
Pharmacogenetic factors determining the metabolism and safety of aromatic anticonvulsants in the residents of Russia
https://doi.org/10.47183/mes.2025-364
Abstract
Introduction. The use of aromatic anticonvulsants (carbamazepine, oxcarbazepine, lamotrigine, and phenytoin) is associated with the risk of severe hypersensitivity reactions, which are partly dependent on HLA-B and HLA-A genotypes (for carbamazepine/oxcarbazepine). Phenytoin and fosphenytoin are metabolized by CYP2C9; therefore, in the setting of genetically determined impaired drug tolerability, the likelihood of adverse events increases. Consideration of the CYP2C9 genotype is important for developing personalized drug dosing schemes and improving treatment outcomes.
Objective. Assessment of the prevalence of major pharmacogenetic variants associated with response to aromatic anticonvulsants, with geographic stratification and identification of at-risk populations warranting preemptive genotyping prior to treatment initiation.
Materials and methods. The study was performed using samples from the Population Frequency Database (GDB) of the Federal Medical and Biological Agency (FMBA) of Russia (n = 120,979, covering 82 RF subjects). Whole-genome sequencing of DNA samples was conducted followed by an analysis of the carrier frequency of HLA-B*15:02, HLA-B*15:11, HLA-A*31:01, and various allelic variants of CYP2C9 with calculation of the enzyme activity score. These metrics were compared across different Russian regions, identifying high-risk biogeographic groups.
Results. The HLA-B*15:02 variant showed a prevalence of less than 1% in all regions of the Russian Federation. A relatively high carrier frequency of HLA-B*15:11 was observed in the Republics of Buryatia and Tyva (1.3%, p = 7.7 × 10–5 and 3.46%, p = 2.4 × 10–3, respectively, compared to a population frequency of 0.11%). The elevated frequencies of HLA-A*31:01 were detected in Perm Krai and the Republics of Kalmykia, Buryatia, Tyva, and Sakha (Yakutia) (8.48%, p = 0.042; 8.79%, p = 0.044; 10.3%, p = 3.4 × 10–10; 20.44%, p = 3.4 × 10–10; 28.74%, p = 5.4 × 10–122, respectively, compared to a population frequency of 5.06%). The Republics of Dagestan, Ingushetia, and Kabardino-Balkaria showed a higher prevalence of impaired metabolism phenotype for phenytoin/phosphenytoin (46.4%, p = 5.6 × 10–36; 44.69%, p = 1.7 × 10–13; 43.83%, p = 1.9 × 10–16), primarily due to a high frequency of the CYP2C9*3 allele. The Republics of Tatarstan, Mari El, and Chuvashia were also characterized by a high incidence of alleles associated with impaired metabolism of these drugs (37.06%, p = 0.028; 37.99%, p = 0.031; 41.2%, p = 5.3 × 10–10), attributable to the presence of the generally rare CYP2C9*29 allele in their genetic structure.
Conclusions. The results obtained enable the formulation of region-specific recommendations for personalizing treatment with aromatic anticonvulsants. For residents of Sakha (Yakutia), Tyva, Buryatia, Kalmykia, and Perm Krai, testing for HLA-A*31:01 carriage is justified. For residents of Tyva and Buryatia, additional testing for HLA-B*15:11 carriage is warranted prior to the prescription of carbamazepine and oxcarbazepine. Before initiating phenytoin therapy, CYP2C9 genotyping is particularly important for the populations of Dagestan, Ingushetia, Kabardino-Balkaria, Tatarstan, Mari El, and Chuvashia. However, this genotyping can be recommended for the entire population of Russia due to the high prevalence of alleles associated with reduced and absent enzyme activity.
Keywords
For citations:
Spektor E.D., Yudin V.S., Mamchur A.A., Rumyantseva A.M., Ivanov M.V., Mitrofanov S.I., Snigir E.A., Keskinov A.A., Yudin S.M., Kashtanova D.A. Pharmacogenetic factors determining the metabolism and safety of aromatic anticonvulsants in the residents of Russia. Extreme Medicine. 2026;28(1):51-60. https://doi.org/10.47183/mes.2025-364
INTRODUCTION
Antiepileptic drugs (AEDs), or anticonvulsants, are a class of pharmacological agents widely used in neurology and psychiatry. Their application extends beyond seizure control and includes the treatment of migraine,1 cluster headache,2 neuropathic pain,3 management of psychotic and behavioral disorders in elderly patients4 with cognitive impairment, as well as serving as mood stabilizers and anxiolytics [1]. Among these drugs, carbamazepine, oxcarbazepine, lamotrigine, and phenytoin have been extensively studied in terms of pharmacogenetic aspects related to their safety profiles. These drugs share a common structural feature — a benzene ring — leading to their classification as “aromatic anticonvulsants” in the English-language literature [2]. The use of this drug class is associated with a low, yet clinically significant, risk of hypersensitivity reactions, such as severe cutaneous adverse reactions (SCARs), including epidermolytic drug reactions (e.g., Stevens–Johnson Syndrome (SJS), toxic epidermal necrolysis (TEN)), and systemic drug reactions with multiorgan involvement (e.g., Drug Reaction with Eosinophilia and Systemic Symptoms (DRESS)).
SJS and TEN are acute, severe allergic reactions characterized by extensive lesions of the skin and mucous membranes. They are currently viewed as different stages of the same pathological process, rather than as separate diseases [3, 4]. The incidence of SJS is 2.5–9.2 cases per million per year with a mortality rate of 4.1–16.9%. For TEN, the incidence amounts to 0.94–1.9 cases per million per year with a mortality rate of 14.8–29.9% [4]. The DRESS syndrome is characterized by widespread (> 50% of body surface area) exanthematous rash, fever, lymphadenopathy, eosinophilia, and internal organ involvement [5]. The exact incidence of DRESS is unknown, being estimated at one case per 10,000 drug exposures in the treated population. According to various estimates, the mortality rate ranges from 3.8% to 10–20% [6][7]. Lamotrigine is among the top five drugs most strongly associated with fatal outcomes in the DRESS syndrome [8]. Furthermore, it ranks second among the medications most frequently implicated in the development of this syndrome [1].
Genetic predisposition plays a significant role in the likelihood of developing the aforementioned allergic reactions. Such predisposition is associated with the carriage of specific alleles of human leukocyte antigens A and B (HLA-A, HLA-B). These genes are part of a larger cluster known as the major histocompatibility complex. Accumulated evidence indicates the association between the development of SCAR/DRESS upon administration of phenytoin, lamotrigine, carbamazepine, and oxcarbazepine with the carriage of HLA-B*15:02 (Level 1A) [9][10]. Furthermore, the association between carbamazepine-induced SCAR/DRESS and the carriage of HLA-A*31:01 (Level 1A), HLA-B*15:11 (Level 1A), and HLA-B*40:01 (Level 2A) was also confirmed [10][11].
Currently, two primary sources provide pharmacogenetic recommendations to guide the prescription of aromatic anticonvulsants. These are the guidelines from the Clinical Pharmacogenetics Implementation Consortium (CPIC) [9][11] and those from the Dutch Pharmacogenetics Working Group (DPWG) [10]. According to CPIC, carbamazepine, oxcarbazepine, or phenytoin should be avoided in carbamazepine-naive patients who are carriers of the HLA-B*15:02 allele. However, in cases where a patient has been using the target drug for three months or longer without developing cutaneous adverse reactions, the therapy may be continued regardless of HLA-B genotype. Indeed, SCARs typically manifest within the first 4–28 days of treatment [9][11]. A similar decision-making algorithm is recommended for the prescription of carbamazepine in carriers of the HLA-A*31:01 allele: avoid prescribing to carriers, except in situations where carbamazepine has been used continuously for three months or more without adverse effects [11].
The DPWG guidelines encompass a broader range of clinical scenarios. According to these recommendations, the use of lamotrigine is contraindicated in carriers of the HLA-B*15:02 allele. Furthermore, the administration of carbamazepine is not recommended for carriers of the HLA-B*15:02 and HLA-A*31:01 alleles (also indicated by CPIC), as well as for carriers of the HLA-B*15:11 allele [10].
Phenytoin and its less commonly used prodrug — fosphenytoin — possess a narrow therapeutic index. Their tolerability profile, in addition to the hypersensitivity reactions discussed above, includes a wide spectrum of adverse effects, such as central nervous system toxicity, nausea, vomiting, gingival hyperplasia, osteomalacia, hematological suppression, and cognitive impairment [9]. The risk of developing these effects is largely determined by the rate of drug biotransformation, which is primarily mediated by the cytochrome P450 isoenzyme CYP2C9.
Numerous allelic variants of the CYP2C9 gene exist, each conferring a predictable activity level of the encoded enzyme. The Activity Score (AS) system is used to describe the functional impact of a single allele (haplotype), classifying haplotypes as having normal (AS = 1), decreased (AS = 0.5), or zero (AS = 0) activity. Allele annotations are provided in the PharmGKB database [12]. The most common variants are CYP2C9*1 (AS = 1), CYP2C9*2 (AS = 0.5), and CYP2C9*3 (AS = 0).
The clinically significant metabolic phenotype is determined based on the patient’s diploid genotype, i.e., the sum of the AS values from both inherited alleles. According to international guidelines, this total score allows for classification into the following phenotypic groups: normal metabolizer (AS = 2), intermediate metabolizer (1 ≤ AS < 2), and poor metabolizer (AS < 1). CPIC guidelines specify maintenance dose adjustments based on the metabolic phenotype: a 25% reduction of the standard maintenance dose is recommended for intermediate metabolizers with AS = 1, and a 50% reduction for poor metabolizers, while the loading dose regimen remains unchanged [9].
Therefore, when prescribing aromatic antiepileptic drugs, the pharmacogenetic profiles of patients should be taken into account. This enables risk management for serious hypersensitivity-mediated adverse reactions, prediction of individual drug metabolism rates, and development of personalized dosing regimens to improve treatment tolerability and its outcomes.
In this study, we aim to assess the population prevalence of major pharmacogenetic variants associated with response to aromatic antiepileptic drugs and to carry out their geographic stratification to identify high-risk populations where preemptive genotyping is clinically justified prior to initiating this drug class.
MATERIALS AND METHODS
Study object
The study was conducted using a population-based sample from the Population Frequency Database (GDB) of the Centre for Strategic Planning and Management of Biomedical Health Risks of the FMBA.5 The sample size comprised 120,979 individuals (58,339 (48.2%) males, mean age 50.1 ± 14.1 years (M ± SD)) from 82 federal subjects of the Russian Federation. For the analysis of regional allele frequency distribution, participants from the city of Moscow and Moscow Oblast, the city of Saint Petersburg and Leningrad Oblast, and the city of Sevastopol and the Republic of Crimea were pooled. The recruitment criteria for the GDB participants can be found in a previously published work [13]. For comparative analysis, regions with a sample size of 250 individuals or more were selected.
DNA extraction and sequencing
Genomic DNA was extracted from whole blood samples using MagAttract HMW DNA Kits (Qiagen, Germany) for high-molecular-weight DNA isolation with magnetic particles on automated liquid handling workstations. Whole-genome sequencing libraries were prepared using an Illumina DNA Prep reagent kit according to the manufacturer’s protocol (Document No. 1000000025416 v10, Illumina, USA) with IDT-ILMN Nextera DNA UD Indexes Set A and Set B, followed by quality control. Whole-genome sequencing was performed on the Illumina NovaSeq 6000 platform using an S4 Reagent Kit (300 cycles) (Illumina, USA) for paired-end reads of 2 × 150 bp.
During the demultiplexing stage, sequencing data in BCL format were converted to FASTQ format using Illumina bcl2fastq2 v2.20.0.422.6 Quality control of the sequencing run was performed with Illumina Sequencing Analysis Viewer v2.4.7,7 while read quality (FASTQ.GZ format) was assessed using the FastQC v0.11.9 bioinformatic tool.8
Reads were aligned to the GRCh38.d1.vd19 reference genome using the Illumina DRAGEN Bio-IT Platform v07.021.510.3.5.7.10 Quality control of the alignments (BAM format) was performed using DRAGEN, FastQC v0.11.9,11 samtools v1.13 [14], and mosdepth v0.3.1 [15]. All samples passed quality control based on key metrics, including the percentage of duplicated reads, the number of unaligned reads, etc. The final mean genome coverage per sample was at least 30x.
Detection of small genetic variants (up to 50 bp) was performed using Illumina Strelka2 v2.9.10 [16]. The sample set was checked for duplicates using Picard CrosscheckFingerprints [17] with the built-in map file. The bioinformatic processing pipeline used was validated (F-score = 99.83%) against the HG001 reference genome provided by the Genome In A Bottle (GIAB) consortium (v.3.3.2) [18].
Pharmacogenetic variant screening and data statistical analysis
Selection of pharmacogenetic variants for analysis was based on their inclusion in existing clinical pharmacogenetic guidelines (CPIC or DPWG). The following allelic variants met this criterion: human leukocyte antigen alleles HLA-B*15:02, HLA-B*15:11, HLA-A*31:01, and variants of the cytochrome P450 isoenzyme gene CYP2C9. CYP2C9 haplotyping was performed using the publicly available Panno algorithm.12 Unphased HLA typing for six major loci was conducted using the xHLA [ 04.10.2017] (utilizing two IPD-IMGT/HLA database versions: 3.22.0 [ 10.2015] and 3.45.1 [ 08.2021]) and HLAscan v2.1.4 [ 04.12.2019] (IPD-IMGT/HLA database version 3.21.0 [ 07.2015]) bioinformatic tools.
For each allele, the carrier frequency and 95% confidence interval (CI), calculated using the Wilson method, were determined. Based on the CYP2C9 allele annotations provided in PharmGKB [13], a CYP2C9 Activity Score (AS) was calculated for each sample. The corresponding metabolic phenotype for enzyme substrates was subsequently assigned based on the AS according to established algorithms. The proportion of individuals in the population requiring personalized phenytoin/fosphenytoin dosing per CPIC guidelines [9] was calculated, along with the combined proportion of intermediate and poor metabolizers, subsequently designated as the “impaired metabolism” group.
Statistical analysis was performed using the Python programming language, version 3.9. A z-test for proportions was used to compare the carrier frequencies of HLA alleles and the proportions of CYP2C9 substrate impaired metabolizers between regions and the overall population values. The Benjamini–Hochberg procedure, controlling the False Discovery Rate (FDR) at 5%, was applied to correct for multiple testing. Associations were considered statistically significant either at a p-value < 0.05 or by the interval method when the confidence interval did not cross a predefined critical threshold.
RESULTS
Severe risk of hypersensitivity reaction in the population treated with aromatic antiepileptic drugs
The absolute counts and percentage frequencies of the risk alleles HLA-B*15:02, HLA-B*15:11, and HLA-A*31:01 were 22 (0.01%), 134 (0.06%), and 6122 (2.53%), respectively. The estimated carrier frequencies for these alleles were 0.02% (HLA-B*15:02), 0.11% (HLA-B*15:11), and 5.06% (HLA-A*31:01). The presence of at least one risk allele was observed in 5.18% of the population. This was primarily driven by the HLA-A*31:01 allele, since the other alleles are extremely rare in the general population. Analysis of the regional distribution of risk allele carriers revealed regions with both statistically significantly lower and higher carrier frequencies compared to the overall population. Table 1 presents data for regions where the frequency of risk alleles for SCAR/DRESS was significantly higher than the population average.
Table 1. High-prevalence regions for risk alleles causing drug-induced epidermolysis and DRESS syndrome
|
Allelic variant |
Carrier status in the population, % |
Regions with higher prevalence |
Carrier status in the region, % |
p-adj |
|
HLA-B*15:02 |
0.02 |
no region |
N/A |
N/A |
|
HLA-B*15:11 |
0.11 |
Buryatia |
1.3% |
7.7×10-5 |
|
Tyva |
3.46% |
2.4×10-3 |
||
|
Sakha (Yakutia) |
0.68% |
3.2×10-3 |
||
|
HLA-A*31:01 |
5.06 |
Perm Krai |
8.48% |
0.042 |
|
Kalmykia |
8.79% |
0.044 |
||
|
Buryatia |
10.3% |
3.4×10-10 |
||
|
Tyva |
20.44% |
3.4×10-10 |
||
|
Sakha (Yakutia) |
28.74% |
5.4×10-122 |
Table compiled by the authors based on original data
Note: p-adj — FDR-corrected p-value (Benjamini–Hochberg false discovery rate); N/A — not applicable.
The HLA-B*15:02 allele exhibits an extremely low prevalence in the general population and was not detected in 53 out of 68 regions. The highest frequency nationwide was observed in Kalmykia (0.52%, p-adj = 0.21).
For the HLA-B*15:11 and HLA-A*31:01 alleles, distinct high-risk regions can be identified, which partially overlap (Table 1). The HLA-B*15:11 variant was not detected in 33 regions. A regional risk assessment for carrying at least one of these risk alleles is illustrated in Figure 1. Based on interval estimation, the republics of Sakha (Yakutia), Tyva, Buryatia, Kalmykia, and Perm Krai demonstrate a statistically significant elevated risk of carriage for the HLA-B*15:02, HLA-B*15:11, or HLA-A*31:01 alleles.
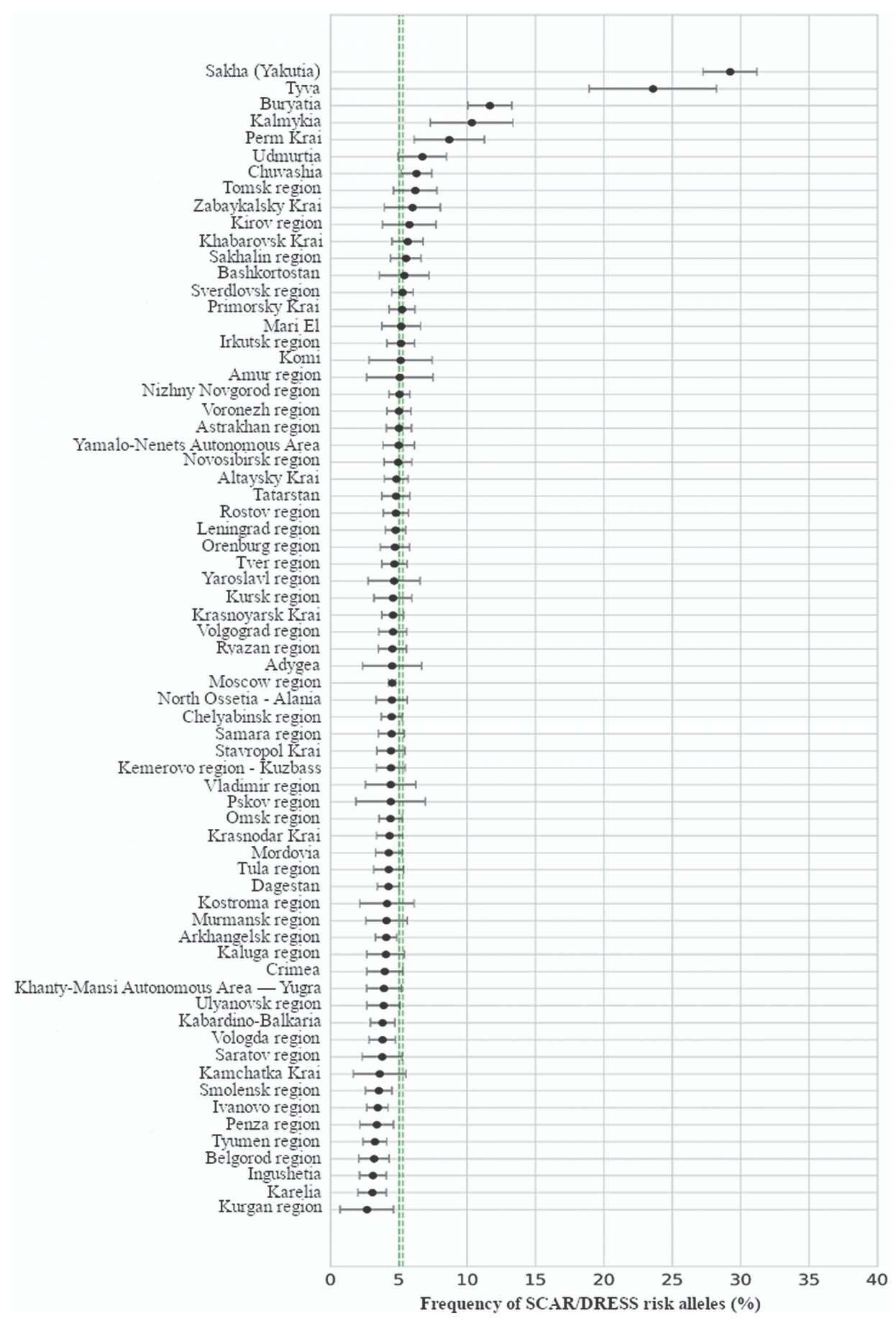
Figure prepared by the authors based on original data
Fig. 1. Allele frequency of risk variants (HLA-B*15:02, HLA-B*15:11, or HLA-A*31:01 combined) in regions of the Russian Federation: the double dashed line represents the 95% confidence interval (CI) for the population-wide prevalence
Phenytoin metabolism in the population
CYP2C9 genotype data were available for 119,422 participants (48.4% males). Figure 2 presents the frequency distributions of the most common alleles, enzyme activity scores, and metabolic phenotypes in the population. The *1 (reference), *2, and *3 allelic variants account for 99% of all detected alleles (Fig. 2A).

Figure prepared by the authors based on original data
Fig. 2. Population-Wide Characterization of CYP2C9 Genetics: A — prevalence of the most frequent alleles; B — distribution of the genotype-predicted activity score; C — distribution of metabolic phenotypes
Approximately one-third (33.6%) of the population exhibits impaired (intermediate or poor) metabolism of CYP2C9 substrates, although the poor metabolizer phenotype is relatively rare (2% of cases). Table 2 presents data for regions with statistically significant higher or lower prevalence of impaired metabolizers of CYP2C9 substrates.
Table 2. Comparative analysis of the prevalence of different CYP2C9 substrate metabolism phenotypes in regions with deviating frequencies of slow metabolizers (intermediate + poor) from the general population
|
Direction of differences between regional and population estimates |
Region |
Prevalence, % |
p-adj |
||
|
Prevalence of impaired metabolism (intermediate + poor) |
Prevalence of intermediate metabolism phenotype (AS 1–1.5) |
Prevalence of poor metabolizer phenotype (AS 0–0.5) |
|||
|
The proportion of impaired metabolizers is lower than the population value |
Sakha (Yakutia) |
17.6 |
17.01 |
0.59 |
2.7×10-78 |
|
Tyva |
17.98 |
17.35 |
0.63 |
5.9×10-12 |
|
|
Kalmykia |
21.3 |
19.22 |
2.08 |
3.6×10-8 |
|
|
Buryatia |
21.98 |
21.2 |
0.78 |
1.7×10-26 |
|
|
Kemerovo region |
30.11 |
28.57 |
1.54 |
0.031 |
|
|
The proportion of impaired metabolizers is higher than the population average |
Tatarstan |
37.06 |
34.56 |
2.5 |
0.028 |
|
Mari-El |
37.99 |
34.6 |
3.39 |
0.031 |
|
|
Chuvashia |
41.2 |
38.06 |
3.14 |
5.3×10-10 |
|
|
Kabardino-Balkaria |
43.83 |
39.32 |
4.51 |
1.9×10-16 |
|
|
Ingushetia |
44.69 |
38.53 |
6.16 |
1.7×10-13 |
|
|
Dagestan |
46.4 |
39.64 |
6.76 |
5.6×10-36 |
|
Table compiled by the authors based on original data
Note: p-adj — FDR-adjusted p-value (Benjamini-Hochberg false discovery rate) for comparison of the total proportion of impaired metabolizers (intermediate + poor) with the population value.
Several republics in the North Caucasus (Kabardino-Balkaria, Ingushetia, and Dagestan), along with the republics of Tatarstan, Mari El, and Chuvashia, can be characterized as high-risk regions for poor tolerability during treatment with phenytoin/fosphenytoin, as well as other CYP2C9 substrate medications not examined in the present study. Conversely, in Sakha (Yakutia), Tyva, Kalmykia, Buryatia, and the Kemerovo region, reduced CYP2C9 enzyme activity is relatively uncommon.
To elucidate the underlying causes of the differences between high-risk regions and the general population, we analyzed the structure of CYP2C9 allelic variants. The observed disparities could be attributed either to an increased frequency of the *2 and *3 alleles or to the presence of region-specific rare alleles. Variants with a prevalence exceeding 1% in any given region are presented in Table 3 alongside the corresponding population-wide frequencies.
Table 3. Frequency of CYP2C9 allelic variants in regions with elevated prevalence of genotypes associated with impaired metabolism of enzyme substrates
|
Region |
n |
Allelic variant |
Regional frequency, % |
Population frequency, % |
Statistical significance level, р |
|
Tatarstan |
1600 |
*2 |
9.53 |
9.62 |
0.87 |
|
*3 |
8.59 |
7.7 |
0.17 |
||
|
*29 |
2.16 |
0.99 |
0.002 |
||
|
Mari-El |
954 |
*2 |
7.94 |
9.62 |
0.074 |
|
*3 |
7.94 |
7.7 |
0.74 |
||
|
*29 |
5.6 |
0.99 |
4.2×10-9 |
||
|
Chuvashia |
1779 |
*2 |
7.81 |
9.62 |
0.005 |
|
*3 |
8.89 |
7.7 |
0.07 |
||
|
*29 |
6.32 |
0.99 |
1.8×10-20 |
||
|
Kabardino-Balkaria |
1709 |
*2 |
11.43 |
9.62 |
0.02 |
|
*3 |
12.7 |
7.7 |
3.9×10-10 |
||
|
Ingushetia |
1186 |
*2 |
7.12 |
9.62 |
0.001 |
|
*3 |
18.25 |
7.7 |
4.7×10-21 |
||
|
Dagestan |
2470 |
*2 |
8.52 |
9.62 |
0.056 |
|
*3 |
17.87 |
7.7 |
3.4×10-40 |
Table compiled by the authors based on original data
Note: the presented alleles have a regional frequency of > 1%; N — number of observations in the region; statistically significant differences (p < 0.05) are highlighted in bold.
Thus, the increased frequency of genotypes associated with impaired metabolism of CYP2C9 substrates in Kabardino-Balkaria, Ingushetia, and Dagestan is driven by a higher prevalence of the CYP2C9*3 variant, which corresponds to severely reduced enzyme activity. A different pattern is observed in Tatarstan, Mari El, and Chuvashia. Here, while the frequencies of the *2 and *3 alleles are comparable to the general population, the primary contributing factor is the relatively high prevalence of the *29 allele, which also confers reduced enzyme activity.
DISCUSSION
Our results have enabled the identification of both population-wide trends and risks associated with the use of AEDs, as well as the stratification of the population to delineate biogeographic groups requiring particular caution when prescribing aromatic anticonvulsants.
Overall, the population of the Russian Federation can be characterized as having a low risk of developing severe toxic-allergic reactions, such as Stevens–Johnson syndrome, toxic epidermal necrolysis, and DRESS syndrome, when using aromatic AEDs. However, a moderate risk of an intermediate phenytoin metabolism phenotype has been established, necessitating a corresponding 25% reduction in the drug maintenance dose [9]. Failure to account for this factor may not only worsen treatment tolerability and increase temporal and financial costs associated with switching AEDs, but also lead to the development of drug resistance [19].
The populations in such regions as Sakha (Yakutia), Tyva, Buryatia, Kalmykia, and to a lesser extent Perm Krai, can be identified as high-risk populations for carrying allelic variants associated with severe hypersensitivity reactions to carbamazepine and oxcarbazepine. This is due to the relatively high prevalence of HLA-B*15:11 and HLA-A*31:01 variants. However, this conclusion cannot be extended to lamotrigine and phenytoin, since the only reliably established association for hypersensitivity reactions to these drugs currently exists with the HLA-B*15:02 allele. This allele has a frequency of less than 1% across all biogeographic groups in Russia.
According to the literature, HLA-B*15:02 and HLA-A*31:01 alleles are globally rare, yet their carrier frequency exhibits ancestry dependence. An elevated frequency of the HLA-B*15:02 variant has been described in indigenous populations of East Asia (6.9%), Australia and Oceania (5.4%), and South/Central Asia (4.6%). Similarly, the HLA-A*31:01 variant is more prevalent in individuals of European ancestry (3%), Hispanic populations in South America (6%), and some East Asian populations (up to 8%) [11].
Notably, the high-risk regions for SCAR/DRESS identified in our study are heterogeneous in terms of the specific alleles driving this risk. For the prevention of severe hypersensitivity reactions to carbamazepine/oxcarbazepine, preemptive testing for HLA-A*31:01 carriage appears sufficient in the republics of Sakha (Yakutia), Kalmykia, and Perm Krai. In contrast, in the republics of Tyva and Buryatia, testing for both HLA-B*15:11 and HLA-A*31:01 is warranted, with subsequent clinical decisions guided by current CPIC guidelines [11]. However, precise determination of the optimal diagnostic panel requires further health-economic analysis.
The structure of CYP2C9 allelic variants in the Russian population highlights its distinct characteristics. On the one hand, the leading allelic variants, aside from the wild-type (*1), are *2 and *3, which aligns with the global pattern and are included in the recommended diagnostic genotyping panel for this enzyme [21]. On the other hand, the frequency of CYP2C9*2 (0.096) is somewhat higher than in the European population (0.076), most closely resembling the corresponding value in Middle Eastern populations (0.083) [12][21].
Concurrently, nearly 2% of the Russian population are carriers of the CYP2C9*29 allele, which is exceptionally rare across all biogeographic groups [21]. This allele encodes an enzyme with reduced activity (AS = 0.5) yet is not included in the extended diagnostic panel recommended by international guidelines [20].
The analysis of the prevalence of genetically determined impaired phenytoin metabolism has demonstrated that the overall Russian population is characterized by a high prevalence (33.6%) of this phenotype. This necessitates consideration not only when prescribing phenytoin/fosphenytoin but also for warfarin [22], a number of non-steroidal anti-inflammatory drugs [24], and other medications whose biotransformation involves the CYP2C9 isoenzyme.
The issue of reduced CYP2C9 enzyme activity is particularly relevant in the Republics of Tatarstan, Mari El, Chuvashia, Kabardino-Balkaria, Ingushetia, and Dagestan, where the frequency of genotypes conferring reduced CYP2C9 activity exceeds the population average, reaching 37–46%. However, the underlying genetic source differs fundamentally between the group of North Caucasian republics and Tatarstan, Mari El, and Chuvashia.
In the former case, the distribution of common allelic variants is shifted toward a predominance of the CYP2C9*3 allele, which corresponds to severely reduced enzyme activity. Therefore, for residents of Kabardino-Balkaria, Ingushetia, and Dagestan, detection of CYP2C9*2 and CYP2C9*3 alleles can be recommended for personalizing therapy with phenytoin/fosphenytoin and other drugs metabolized by CYP2C9.
In Chuvashia, Tatarstan, and Mari El, the deviation from the population average can be described as qualitative, since the risk of impaired metabolism is driven by a different allele — CYP2C9*29 — which is not widely prevalent in the general population. For this reason, inclusion of this variant in a region-specific diagnostic panel for these areas appears promising. However, similar to HLA typing, the cost-effectiveness of implementing such diagnostic technologies can only be clarified following a formal health-economic analysis.
CONCLUSIONS
Our study demonstrates significant variability in the prevalence of genetic markers associated with impaired metabolism and the risk of adverse reactions to aromatic anticonvulsants among the population of the Russian Federation. While carriage of alleles increasing the risk of SCAR/DRESS is rare in the general population, the high prevalence of some of them (HLA-B*15:11 and HLA-A*31:01) among residents of Sakha (Yakutia), Tyva, Buryatia, Kalmykia, and Perm Krai warrants recommending genetic testing for these variants prior to prescribing carbamazepine and oxcarbazepine in these regions.
The use of lamotrigine appears equally safe across all biogeographic groups of the Russian population due to the low prevalence of the HLA-B*15:02 allele.
Regarding phenytoin, approximately one-third of the population has a genetically determined impaired metabolism of the drug, requiring dose adjustment, particularly in the republics of the North Caucasus, as well as in Tatarstan, Mari El, and Chuvashia. Consequently, before initiating phenytoin therapy, CYP2C9 genotyping is especially important for the population of these regions but can generally be recommended for all residents of Russia.
Authors’ contributions. All authors confirm that they meet the ICMJE criteria for authorship. The largest contribution was distributed as follows: Ekaterina D. Spektor — conceptualization, formal analysis; Vladimir S. Yudin — methodology, manuscript drafting and editing; Aleksandra A. Mamchur — manuscript drafting; Antonina M. Rumyantseva — resources; Mikhail V. Ivanov — software; Sergey I. Mitrofanov — data verification; Ekaterina A. Snigir — conducting research; Anton A. Keskinov — project administration, management of planning and implementation of research activities; Sergey M. Yudin — research management; Daria A. Kashtanova — coordination of planning and implementation of research activities.
1. Migraine. Clinical guidelines. All-Russian Society of Neurologists. Russian Society for the Study of Headache (ROIGB); 2020 (In Russ.).
2. Primary headaches: diagnosis and treatment. Methodological recommendations. Moscow City Health Department; 2017 (In Russ.).
3. Neuropathic pain: clinical guidelines for diagnosis and treatment of the Russian Society for the Study of Pain. Russian Society for the Study of Pain; 2018 (In Russ.).
4. Cognitive disorders in elderly and senile individuals. Russian Society of Psychiatrists; 2020 (In Russ.).
5. Population Genetic Variants Frequency Database of the Russian Federation Population. FMBA of Russia. App version 1.1.3 (17.03.2025). Database version 59.1 (03.10.2024) (In Russ.).
6. bcl2fastq and bcl2fastq2 Conversion Software Downloads. URL: https://emea.support.illumina.com/sequencing/sequencing_software/bcl2fastq-conversion-software/downloads.html
7. Sequencing Analysis Viewer Support. URL: https://support.illumina.com/sequencing/sequencing_software/sequencing_analysis_viewer_sav.html
8. Babraham Bioinformatics — FastQC A Quality Control tool for High Throughput Sequence Data. URL: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
9. GDC Reference Files | NCI Genomic Data Commons. URL: https://gdc.cancer.gov/about-data/gdc-data-processing/gdc-reference-files
10. DRAGEN secondary analysis | Software for NGS data analysis. URL: https://www.illumina.com/products/by-type/informatics-products/dragen-secondary-analysis.html
11. Babraham Bioinformatics — FastQC A Quality Control tool for High Throughput Sequence Data. URL: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
12. PAnno: A Pharmacogenomics Annotation Tool for Clinical Genomic Testing. URL: https://github.com/PreMedKB/PAnno.git
References
1. Żełabowski K, Wojtyshiak K, Ratka Z, Biedka K, Chlopas-Konowalek A. Lamotrigine Therapy: Relation Between Treatment of Bipolar Affective Disorder and Incidence of Stevens–Johnson Syndrome—A Narrative Review of the Existing Literature. Journal of Clinical Medicine. 2025;14(12):4103. https://doi.org/10.3390/jcm14124103
2. Ahmed AF, Sukasem C, Sabbah MA, Musa NF, Noor DAM, Daud NAA. Genetic Determinants in HLA and Cytochrome P450 Genes in the Risk of Aromatic Antiepileptic-Induced Severe Cutaneous Adverse Reactions. Journal of Personalized Medicine. 2021;11(5):383. https://doi.org/10.3390/jpm11050383
3. Hasegawa A, Abe R. Recent advances in managing and understanding Stevens–Johnson syndrome and toxic epidermal necrolysis. F1000Research. 2020;9:F1000 Faculty Rev-612. https://doi.org/10.12688/f1000research.24748.1
4. Watanabe Y, Hama N. Recent advances in the diagnosis and treatment of Stevens–Johnson syndrome/toxic epidermal necrolysis. Allergology International. 2025;74(3):345–55. https://doi.org/10.1016/j.alit.2025.05.008
5. Mori F, Caffarelli C, Caimmi S, Bottau P, Liotti L, Franceschini F, et al. Drug reaction with eosinophilia and systemic symptoms (DRESS) in children. Acta Biomedica. 2019;90(3-S):66–79.
6. Calle AM, Aguirre N, Ardila JC, Villa RC. DRESS syndrome: A literature review and treatment algorithm. World Allergy Organization Journal. 2023;16(3):100673. https://doi.org/10.1016/j.waojou.2022.100673
7. López-Rocha E, Blancas L, Rodriguez-Mireles K, Gaspar-Lopez A, O’Farrill-Romanillos P, Amaya-Mejia A, et al. Prevalence of DRESS syndrome. Revista Alergia Mexico. 2014;61(1):14–23.
8. Liang C, An P, Zhang Y, Liu X, Zhang B. Fatal outcome related to drug reaction with eosinophilia and systemic symptoms: a disproportionality analysis of FAERS database and a systematic review of cases. Frontiers in Immunology. 2024;15:1490334. https://doi.org/10.3389/fimmu.2024.1490334
9. Karnes JH, Rettie AE, Somogyi AA, Huddart R, Fohner AE, Formea CM, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for CYP2C9 and HLA-B Genotypes and Phenytoin Dosing: 2020 Update. Clinical Pharmacology and Therapeutics. 2021;109(2):302–9. https://doi.org/10.1002/cpt.2008
10. Manson LEN, Nijenhuis M, Soree B, de Boer-Veger NJ, Buunk AM, Houwink EJF, et al. Dutch Pharmacogenetics Working Group (DPWG) guideline for the gene-drug interaction of CYP2C9, HLA-A and HLA-B with anti-epileptic drugs. European Journal of Human Genetics. 2024;32(8):903–11. https://doi.org/10.1038/s41431-024-01572-4
11. Phillips EJ, Sukasem C, Whirl-Carrillo M, Muller DJ, Dunnenberger HM, Chantratita W, et al. Clinical Pharmacogenetics Implementation Consortium Guideline for HLA Genotype and Use of Carbamazepine and Oxcarbazepine: 2017 Update. Clinical Pharmacology and Therapeutics. 2018;103(4):574–81. https://doi.org/10.1002/cpt.1004
12. Barbarino JM, Whirl-Carrillo M, Altman RB, Klein TE. PharmGKB: A worldwide resource for pharmacogenomic information. WIREs. 2018;10(4):e1417. https://doi.org/10.1002/wsbm.1417
13. Gusakova M, Dzhumaniiazova I, Zelenova E, Kashtanova D, Ivanov M, Mamchur A, et al. Prevalence of the cancer-associated germline variants in Russian adults and long-living individuals: using the ACMG recommendations and computational interpreters for pathogenicity assessment. Frontiers in Oncology. 2024;14:1420176. https://doi.org/10.3389/fonc.2024.1420176
14. Li H, Handsaker B, Wysoker A, Fennel T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25(16):2078. https://doi.org/10.1093/bioinformatics/btp352
15. Pedersen BS, Quinlan AR. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics. 2018;34(5):867–8. https://doi.org/10.1093/bioinformatics/btx699
16. Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, et al. Strelka2: fast and accurate calling of germline and somatic variants. Nature Methods. 2018;15(8):591–4. https://doi.org/10.1038/s41592-018-0051-x
17. Najafov J, Najafov A. CrossCheck: an open-source web tool for high-throughput screen data analysis. Scientific Reports. 2017;7(1):5855. https://doi.org/10.1038/s41598-017-05960-3
18. Krusche P, Trigg L, Boutros PC, Mason CE, De La Vega FM, Moore BL, et al. Best practices for benchmarking germline small-variant calls in human genomes. Nature Biotechnology. 2019;37(5):555–60. https://doi.org/10.1038/s41587-019-0054-x
19. Finamore JM, Sperling MR, Zhan T, Nei M, Skidmore CT, Mintzer S. Seizure outcome after switching antiepileptic drugs: A matched, prospective study. Epilepsia. 2016;57(8):1294–300. https://doi.org/10.1111/epi.13435
20. Pratt VM, Cavallari LH, Del Tredici AL, Hachad H, Ji Y, Moyer AM, et al. Recommendations for Clinical CYP2C9 Genotyping Allele Selection: A Joint Recommendation of the Association for Molecular Pathology and College of American Pathologists. Journal of Molecular Diagnostics. 2019;21(5):746–55. https://doi.org/10.1016/j.jmoldx.2019.04.003
21. Huddart R, Fohner AE, Whirl-Carrillo M, Wojcik GL, Gignoux CR, Popejoy AB, et al. Standardized Biogeographic Grouping System for Annotating Populations in Pharmacogenetic Research. Clinical Pharmacology and Therapeutics. 2019;105(5):1256–62. https://doi.org/10.1002/cpt.1322
22. Johnson JA, Caudle KE, Gong L, Whirl-Carrillo M, Stein CM, Scott SA, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for Pharmacogenetics-Guided Warfarin Dosing: 2017 Update. Clinical Pharmacology and Therapeutics. 2017;102(3):397–404. https://doi.org/10.1002/cpt.668
23. Theken KN, Lee CR, Gong L, Caudle KE, Formea CM, Gaedigk A, et al. Clinical Pharmacogenetics Implementation Consortium Guideline (CPIC) for CYP2C9 and Nonsteroidal Anti-Inflammatory Drugs. Clinical Pharmacology and Therapeutics. 2020;108(2):191–200. https://doi.org/10.1002/cpt.1830
About the Authors
E. D. SpektorRussian Federation
Ekaterina D. Spektor, Cand. Sci. (Med.)
Moscow
V. S. Yudin
Russian Federation
Vladimir S. Yudin, Cand. Sci. (Biol.)
Moscow
A. A. Mamchur
Russian Federation
Aleksandra A. Mamchur
Moscow
A. M. Rumyantseva
Russian Federation
Antonina M. Rumyantseva
Moscow
M. V. Ivanov
Russian Federation
Mikhail V. Ivanov
Moscow
S. I. Mitrofanov
Russian Federation
Sergey I. Mitrofanov, Cand. Sci. (Biol.)
Moscow
E. A. Snigir
Russian Federation
Ekaterina A. Snigir, Cand. Sci. (Biol.)
Moscow
A. A. Keskinov
Russian Federation
Anton A. Keskinov, Cand. Sci. (Med.)
Moscow
S. M. Yudin
Russian Federation
Sergey M. Yudin, Dr. Sci. (Med.), Professor
Moscow
D. A. Kashtanova
Russian Federation
Daria A. Kashtanova, Cand. Sci. (Med.)
Moscow
Review
For citations:
Spektor E.D., Yudin V.S., Mamchur A.A., Rumyantseva A.M., Ivanov M.V., Mitrofanov S.I., Snigir E.A., Keskinov A.A., Yudin S.M., Kashtanova D.A. Pharmacogenetic factors determining the metabolism and safety of aromatic anticonvulsants in the residents of Russia. Extreme Medicine. 2026;28(1):51-60. https://doi.org/10.47183/mes.2025-364
JATS XML
















